iCOP™
Integrated Cellular 'Omics Platform
The success of ‘Omics and Systems Biology in other areas of biology and medicine has drawn the attention of the bioprocessing industry where their application is increasingly becoming more prevalent. The main area in biomanufacturing where the use of Systems Biology approaches can have major impact is in process optimization and cell engineering. Systems Biology uses the system-level high dimensional data generated via ‘Omics technologies to provide the fundamental knowledge in order to achieve these two goals.
When genomics technologies were first introduced, many users had a hard time deriving benefit from the data generated. Unfortunately, for many, it is even more difficult with Systems Biology. In order to achieve the significant benefits, you must have sophisticated tools and workflows to gather, integrate, mine, and visualize the data so it can be applied to engineer or optimize the system.
Based on these premises ArrayXpress developed iCOP – an integrated platform for bioprocess optimization. iCOP was designed upon the principles of Systems Biology to enable the systematic and directed engineering of production cell lines. Consistent with the QbD philosophy, it is fundamentally a science- and data-driven approach to bioprocessing, allowing scientists and engineers to build in quality from the design stages. The ultimate goal is to enable increased titers, high product quality and predictable and stable bioprocesses, in what we call Next Generation Bioprocessing.
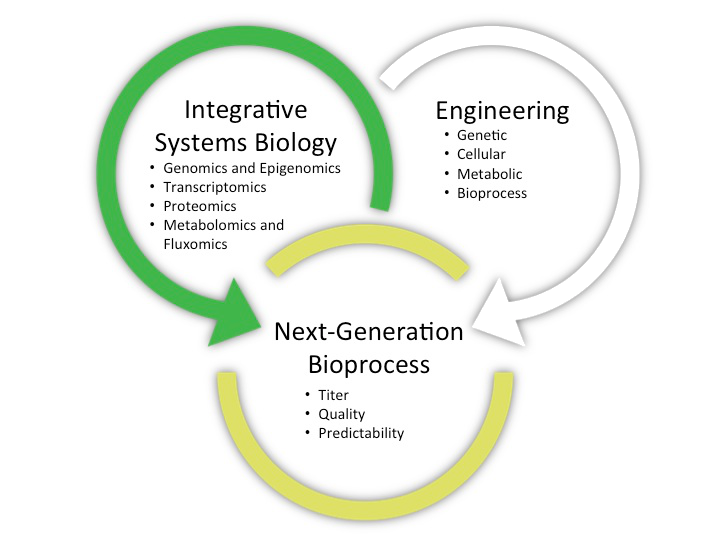
iCOP is comprised of two interconnected arms: an integrative Systems Biology and an Engineering arm. The first relies on system-wide molecular characterization of the cell’s functional dynamics that is used to drive the cell line development and process optimization efforts in the latter.
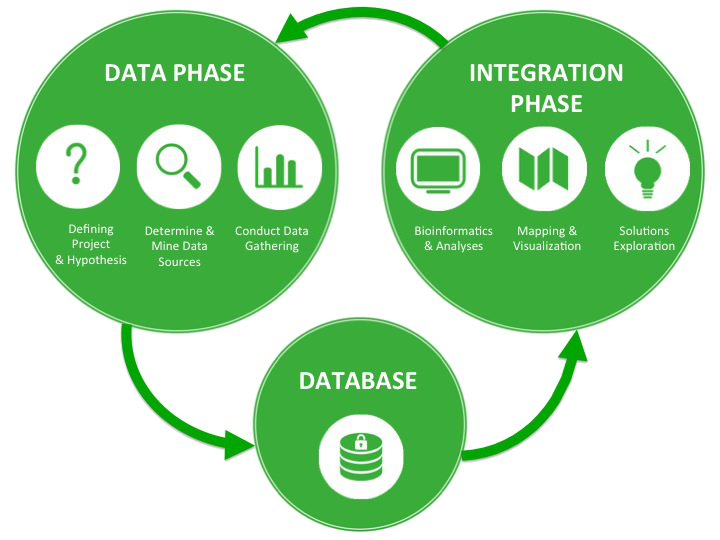
iCOP iterative workflow
The data phase feeds into the proprietary iCOP database which is mined during the integration phase to derive process relevant insights.
iCOP utilizes a comprehensive suite of 'Omics technologies to lay the foundational groundwork to allow for the simultaneous characterization of the various organizational components of the cell: DNA, RNA, proteins, and metabolites. The various 'Omics datasets - genomics, epigenomics, transcriptomics, proteomics, metabolomics, and fluxomics - are integrated to draw insights into the functional dynamics and interactions controlling cellular behavior.
Integration of the various 'Omics datasets and the subsequent interpretation are very demanding, computationally intensive processes that require the highly skilled expertise across various fields in biology and bioinformatics With iCOP, ArrayXpress has developed proprietary data analysis pipelines for the true integration of these varying datasets to derive process relevant insights.
Results of the iCOP Systems Biology Approach
- The creation of an integrated in silico model of the cell that has been tested and refined for robustness and accuracy and that is used for predicting cellular behavior in culture.
- Cellular and process bottlenecks are identified and capitalized for immediate optimization and short-term gains in process improvements.
- Ultimately iCOP's Systems Biology arm enables a knowledge-based system that is capitalized in the Engineering arm.
Simplify Upstream Bioprocess Engineering
- The creation of a collection of engineered cell lines to streamline cell line development and process optimization.
- Optimized cell lines are engineered to achieve a range of critical bioprocess relevant specifications.
- Site-specific integration in a pre-determined genomic location, support high expression levels of transgenes without the need to undergo gene amplification.
- Engineered cellular and metabolic pathways enforce specific product traits required for efficacy and safety based on established Critical Quality Attributes (CQAs).
- Cellular and metabolic engineering across the entire cell line collection maximizes cellular and metabolic outputs.
- Cell line specific biomarkers are used for real-time process monitoring and troubleshooting.
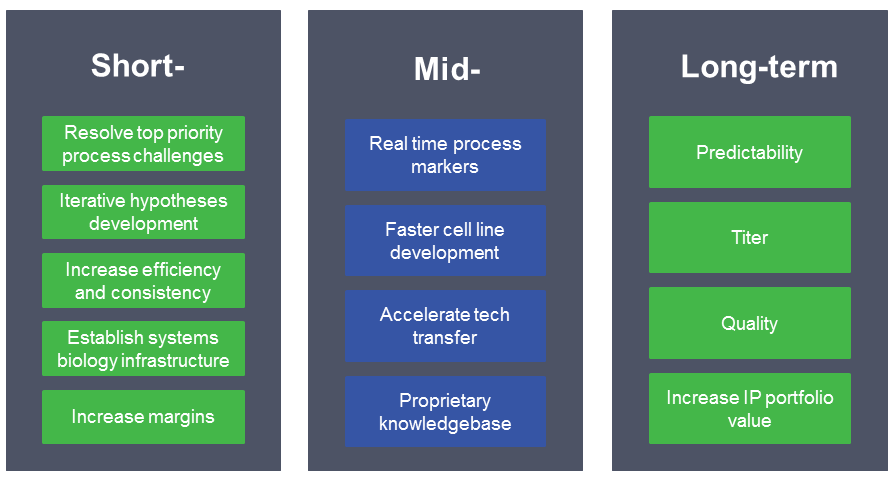
Benefits of iCOP